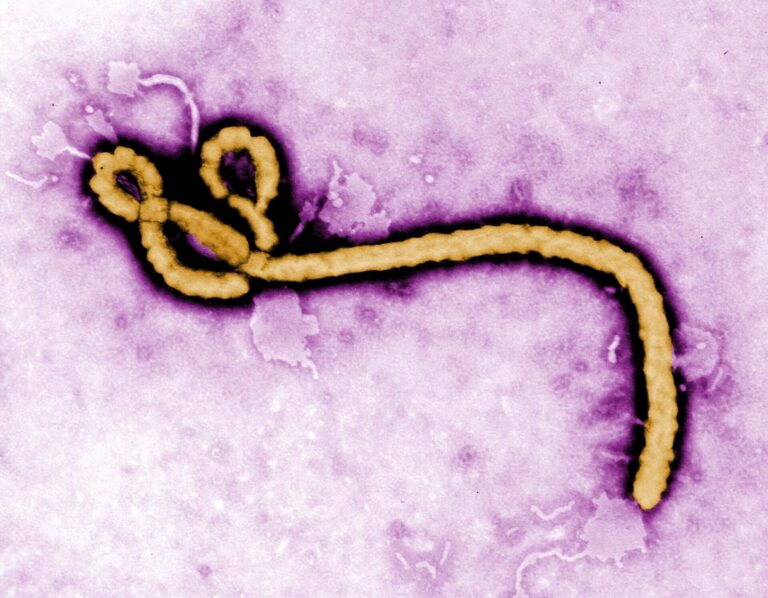
Photo: Thinkstock. The ebola virus, magnified 108,000 times.
Sterghios Moschos, Northumbria University, Newcastle
We have developed technology that reliably detects the Ebola virus in a blood sample seven times faster, ten times cheaper and 700 times safer than doing it in a lab. In fact, the test can be used to detect any genetic material in blood, and so could be used to find everything from malaria to Zika.
Imagine tropical heat. You are wearing a thick, awkward plastic suit that covers you head to toe. Your only access to air is via a respirator; water comes in the form of your own sweat trickling down your eyes. You flex your fingers in your thick, plastic gloves, as you approach yet another person who could kill you: a suffering, feverish, wet patient who may have Ebola virus disease.
You must stick a needle into their vein, get a vial of blood and get it to a diagnostic laboratory. The lab, however, might be several days’ drive away. Until the results are back, the patient, who might have malaria (the symptoms are similar to Ebola), must be kept separate from healthy people. They will be quarantined alongside other suspected cases, some of whom might test positive for Ebola.
Reaching the diagnostic lab, the sample joins a queue of hundreds of samples that need to be manually processed. At the peak of the 2014 West Africa Ebola outbreak, it took five days or more to get a result. By the time the West responded, this was reduced to between five and eight hours. At a cost of US$100 per test, however, the expense remained prohibitive. The average healthcare spend per person per year in oil-rich Nigeria is about US$118, but in Guinea, where the last Ebola outbreak started, it is just US$9. In the UK, it is about US$4,000.
When it comes to Ebola, testing a blood sample for the virus genome is the most reliable diagnostic method. The amount of virus can also tell you if the patient is likely to live or die. However, you first need to get to the genome of the virus and purify it, before the test itself can be applied.
Working on my own diagnostic technologies for lung diseases, I wondered how we could replace this manual, lab-based method with a simple, lab-free one. The goal was to radically cut time and cost, while not making the test less reliable. It was basic science that would lead me and my colleagues to the solution.
Simple, but fit for purpose
The device we developed, the Quantitative, Rapid Identification (QuRapID) platform, uses two simple principles: heat and colour. Freezing breaks up cells and spills their contents. Doing this on a machine (using a cheap Raspberry Pi computer), allows you to get to the genome of the Ebola virus without destroying it in the process.
Next, to measure how many copies of the virus are in a sample, you’d normally use a chemical reaction that produces a green dye – the greener the sample, the more virus is in it. Blood, however, absorbs green light and emits red light, hence its colour. Switching the chemical reaction components from producing a green dye to producing a specific shade of red means that we can now detect increases in that shade of red using a light sensor controlled by the Rapsberry Pi – the more of that shade of red there is in the drop of blood, the more virus there is. Bring the freezing and red dye together and we can measure the virus straight in the drop of blood.
The whole process is just five simple steps, and it takes 70 minutes or less – from pin prick to traffic light answer.

Biogene Ltd, CC BY-NC-SA
It’s not perfect, but it is fit for purpose. While lab-based methods can detect ten times less amounts of virus than the QuRapID platform, the levels of Ebola virus in the blood of patients with symptoms have so far been well above the lower detection limits of the QuRapID. We never got to test this on patients, though; instead, we tested it on cow blood spiked with live Ebola virus. In the 15 months it took us from August 2014 to turn the idea into a fully working instrument – confirmed to work with the live Ebola virus in November 2015 – the West Africa epidemic was practically over.
Tweaking QuRapID to detect other diseases
When another highly infectious disease, the Middle East Respiratory Syndrome (MERS), hit South Korea in August 2015, we calculated that a new test could be produced by public health agencies to use for mass screening within as little as two weeks. At this point, less than 12 months into our Ebola work, we had still not confirmed the QuRapID could do what we wanted. But, in the future, stockpiling instruments and tests for known high-risk diseases, such as Ebola virus disease, would make mass screening capacity available in a matter of days or even hours.
![]() The Ebola virus is not the only threat, however. Some viruses, such as Zika, can cause disease at much lower levels of infection than those of Ebola virus, so the quest is now on to make this genetic testing platform even more sensitive, and to develop tests for other diseases threatening massive outbreaks.
The Ebola virus is not the only threat, however. Some viruses, such as Zika, can cause disease at much lower levels of infection than those of Ebola virus, so the quest is now on to make this genetic testing platform even more sensitive, and to develop tests for other diseases threatening massive outbreaks.
Sterghios Moschos, Associate Professor in Cellular and Molecular Sciences, Northumbria University, Newcastle
This article was originally published on The Conversation. Read the original article.



30 Comments
Pingback: เว็บแทงบอล อันดับ 1 เดิมพันขั้นตํ่า 10 บาท
Pingback: สล็อตเว็บตรง แตกง่าย ไม่มีขั้นต่ำ
Pingback: Bk8
Pingback: เกมไพ่
Pingback: วิเคราะห์บอลวันนี้
Pingback: polaris snowmobiles canada
Pingback: Why bitcoin crashing is much worse than 9/11
Pingback: รับกำจัดปลวก
Pingback: นำเข้าสินค้าจากจีน
Pingback: เทรดทอง
Pingback: slot gacor
Pingback: BIPOC
Pingback: โคมไฟ
Pingback: John Lobb
Pingback: ebet games เปิดให้บริการ เกมคาสิโนออนไลน์ อะไรบ้าง ?
Pingback: พรมรถ
Pingback: Diyala Science
Pingback: Ford Everest
Pingback: Howto-sbobet เว็บรวมเทคนิคแทงบอลขั้นเทพ ปิดแล้ว
Pingback: therich789 เว็บสล็อตน้องใหม่
Pingback: KC9
Pingback: click over here
Pingback: couples massage
Pingback: Aviator game online
Pingback: nagatop situs scam
Pingback: mostbet
Pingback: FORTUNE DRAGON
Pingback: ufabet789
Pingback: fear of god essentials
Pingback: Plinko BGaming